
Localized signaling islands in cells: new targets for precision drugs
Because their range of action is limited to their immediate vicinity, these messaging stations could be ideal targets for precision drug designMedia Contact: Leila Gray 206.685.0381 | leilag@uw.edu
Research results reported this week in the journal Science overturn long-held views on a basic messaging system within living cells.
The findings suggest new approaches to designing precisely targeted drugs for cancer and other serious diseases.

Dr. John D. Scott, professor and chair of pharmacology at the University of Washington School of Medicine and a Howard Hughes Medical Institute Investigator, along with Dr. F. Donelson Smith of the UW and HHMI, led this study, which also involved Drs. Claire and Patrick Eyers and their group at the University of Liverpool.
The researchers explained that key cellular communication machinery is more regionally constrained inside the cell than was previously thought. Communication via this vital system is akin to social networking on your Snapchat account.
Within a cell, the precise positioning of such messaging components allows hormones, the body’s chief chemical communicators, to transmit information to exact places inside the cell. Accurate and very local activation of the enzyme, which Scott and his group studied, helps assure a correct response occurs in the right place and at the right time.
“The inside of a cell is like a crowded city,” said Scott, “It is a place of construction and tearing down, goods being transported and trash being recycled, countless messages, such as the ones we have discovered, keep assembly lines flowing, and packages moving. Strategically switching on signaling enzyme islands allows these biochemical activities to keep the cell alive and is important to protect against the onset of chronic diseases such as diabetes, heart disease and certain cancers.”
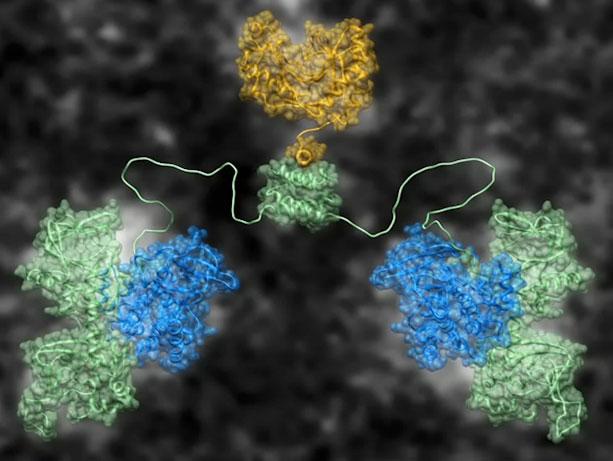
Advances in electron microscopy and native mass spectrometry enabled the researchers to determine that a critical component of the signaling system, anchored protein kinase A, remains intact during activation. Parts of the molecule are flexible, allowing it to both contract and stretch, with floppy arms that can reach out to find appropriate targets.
Still, where the molecule performs its act, space is tight. The distance is, in fact, about the width of two proteins inside the cell.
“We realize that in designing drugs to reach such targets that they will have to work within very narrow confines,” Scott said.
One of his group’s collective goals is figuring out how to deliver precision drugs to the right address within this teeming cytoplasmic metropolis.
“Insulating the signal so that the drug effect can’t happen elsewhere in the cell is an equally important aspect of drug development because it could greatly reduce side effects,” Scott said.
An effort to take this idea of precision medicine a step further is part of the Institute for Targeted Therapeutics at UW Medicine in Seattle. The institute is being set up by Scott and his colleagues in the Department of Pharmacology.

The scientists are collaborating with cancer researchers to better understand the molecular causes -- and possible future treatments -- for a certain liver malignancy. This particular liver cancer arises from a mutation that produces an abnormal form of the enzyme that is the topic of this current work, protein kinase A, and alters the enzyme's role in cell signaling.
Other advances that gave the researchers a clearer view of the signaling mechanisms reported in Science include CRISPR gene editing, live-cell imaging techniques, and more powerful ways to look at all components of a protein complex.
The work was supported by the Howard Hughes Medical Institute, grants from the National Institutes of Health, the Biotechnology and Biological Research Council of the United Kingdom, North West Cancer Research, and the University of Liverpool.
Scott is the Edwin G. Krebs – Speights Professor of Cell Signaling and Cancer Biology at the UW School of Medicine and an investigator of the Howard Hughes Medical Institute.
Grant numbers: 5R01DK105542, 4P01DK05441, 1R01GM120553, BB/L009501/1, and CR1037.
For details about UW Medicine, please visit https://uwmedicine.org/about.