
Embryos unveil their development plans, cell by cell
Studying dynamic changes in chromatin, which stores a cell’s genetic materials, could help unlock how an organism’s many different cell types originally form.Media Contact: EMBL: Iris Kruijen, iris.kruijen@embl.de +49 6221 387 8443
UW Medicine: Leila Gray, leilag@uw.edu, 1 206 685 0381
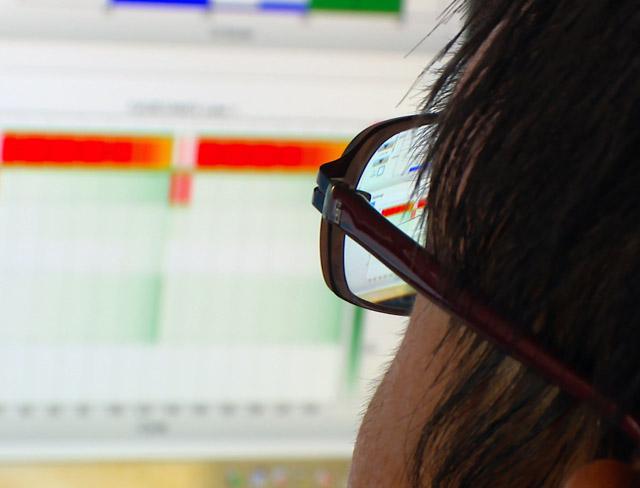

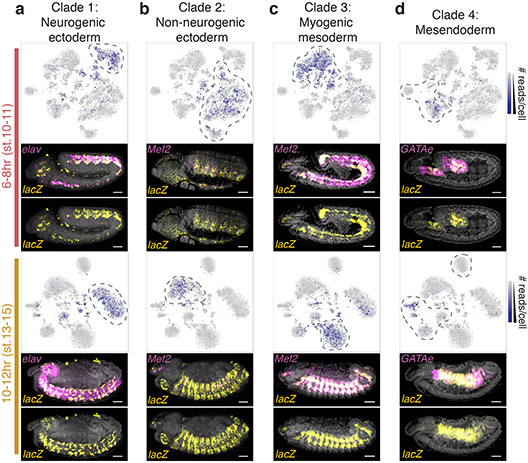
In thousands of single cells from embryos, both cell type and developmental stage can be deduced from measurements of chromatin accessibility.
Chromatin stores the genetic information inside the nucleus of a cells. It is a tightly coiled structure of DNA and proteins. Its dynamic structure can change to open or protect its contents. The usefulness of a new approach to assessing chromatin accessibility in a large number of individual cells is being reported by scientists at the European Molecular Biology Laboratory in Heidelberg, Germany, and UW Medicine in Seattle,
As embryos form and develop, a variety of cells start to appear. The scientists wanted to understand how cells in developing embryos regulate their identity as they determine what kind of cell to become.
Nature published the results of their work on this problem on March 14. Read the paper.
Their new, more systematic approach allows researchers to analyze all the different cell types in an embryo at the same time. They were able to do this at a single-cell resolution.
“I expect this approach to save labs around the world lots of time,” said Eileen Furlong, who co-led the work at EMBL with Jay Shendure, professor of genome sciences at the University of Washington School of Medicine.
Earlier, a researcher would have to first isolate the various cell types, and then investigate the chromatin of each type in separate batches. That lengthy method provided an averaged view across thousands of a given cell type.
“Previous studies have used differences in RNA content to identify cell types and their developmental trajectories,” said Shendure. (RNAs are molecules involved in making proteins and sometimes act as genetic information messengers.)
“Here, we instead measure the state of chromatin in single cells,” he explained. “This contains the regulatory program that governs how and when RNAs are expressed in each cell.
“For the first time, we have looked upstream at how these expression signatures are regulated and therefore drive single cell trajectories during early development,” Furlong added
The chromatin in a human cell contains around two meters of DNA, packed into a nucleus less than one hundredth of a millimeter across. Regulatory elements like promoters and enhancers are short stretches of DNA. They control levels of gene expression and thereby the production of proteins, which are what ultimately make cell types different from one another.
When cells use a particular regulatory element, the chromatin uncoils and its content becomes accessible. That’s why Furlong, Shendure and their research colleagues expected chromatin accessibility to reveals how a cell follows a specific developmental path, such as turning into a highly specialized muscle cell or nerve cell, for example.
Obtaining single-cell information on chromatin accessibility allowed the team to determine a cell’s identity and how that identity is regulated.
They performed the experiments on fruit fly embryos, a model organism for both developmental biology and disease studies. The new approach can be applied to any species.
The results identified thousands of previously unknown regulatory elements that are used in only a subset of cells, and predicted when and where each element is active during development.
The data, made available through a user-friendly browser and an atlas, reveals a wealth of differences between cell types and provides a powerful resource for future studies.
This work was a collaboration between James Reddington and David Garfield in the Furlong group at EMBL, and Darren Cusanovich in Shendure’s lab at the University of Washington School of Medicine. Looking forward, the research team plans to expand this study, and to integrate other layers of single-cell information on the regulation of cell fate decisions during embryogenesis.
EMBL is an intergovernmental European organization established in 1974 and supported by over 20 member states. EMBL performs fundamental research in molecular biology to study the story of life. EMBL has more than 1,600 people, from over 80 countries and operating across six sites in Grenoble, France; Hamburg and Heidelberg in Germany; Cambridge, England, Rome and Barcelona. Its scientists work in independent groups to develop new technology and methods in the life sciences.
For details about UW Medicine, please see our About page.
Topics:cell biology