
Study gives first view of centromere variation and evolution
Critical for accurate transmission of chromosomes during cell division, centromeres differ vastly in size, structure and epigenetic makeup.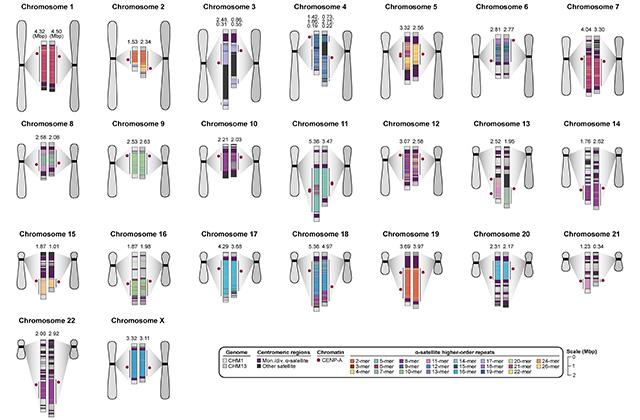
A genomic study of human and selected nonhuman primate centromeres has revealed their unimaginable diversity and speed of evolutionary change.
In cell genetics, a centromere is the spot where two sister chromatids attach. A chromatid is one-half of a duplicated chromosome. United pairs of chromosomes have identifiable shapes because centromeres are not in a uniform position. As a cell prepares to divide, the machinery to separate and segregate chromosomes goes into action at each centromere location.
Unless the genetic material is distributed correctly between the two resulting cells, problems can arise. These include cancer, congenital disorders such as Down syndrome, and the inability of a fertilized cell to grow into a baby.
Although centromeres are vital to proper cell replication, the complexity of their genomic organization had been almost impossible to study. The lack of centromere sequences hindered exploration of how these regions help maintain genetic integrity.
Today, advances in long-read genome sequencing technologies, refined computerized genome assembly algorithms, and improved genome databases are enabling scientists to realize how greatly centromeres differ in size and structure. This opens new avenues toward figuring out what these differences might mean.
A first look at the factors behind the vast variations in centromeres is reported today, April 3, in Nature. The findings suggest that centromeres are likely highly individualized among people. A set of centromeres might even be a personal signature, just as we each have characteristic voice patterns, iris colorations and fingerprints that distinguish us from one another.
It’s not yet known if certain centromere variations might make people susceptible to particular diseases.
Only the human X chromosome appears to be mostly immutable, with very similar sequences and structure across a diversity of humans.
The lead author of the Nature report is Glennis Logsdon, a recent postdoctoral scholar in Evan Eichler’s genomic science lab at the University of Washington School of Medicine in Seattle. Logsdon is now a faculty member at the Perelman School of Medicine at the University of Pennsylvania. Eichler, who is also a Howard Hughes Medical Institute Investigator and a member of the Brotman Baty Institute for Precision Medicine, is the senior and corresponding author.
“This recent research is a direct application of both the Human Pangenome Reference Consortium and the Telomere-to-Telomere (T2T) sequencing efforts to provide new biological insights into complex regions of the human genome critical for chromosome segregation,” noted Eichler. He is known for his work on genome evolution and its potential to create new functions and also for studies of genetic instability associated with disease.
To capture information on how human centromeres might have evolved, the team compared human genetic sequences of two completely sequenced human centromeres with those of some nonhuman primates. These were the chimpanzee and the orangutan, which are great apes and closely related species to humans, and the macaque, an Old World monkey and a more distant relative.
The scientists discovered that centromeres are evolving much faster than other unique portions of the human genome. They are among the most mutation-prone regions of the human genome. The researchers also found that the unique sequences and structure of centromeres were the culmination of different evolutionary forces moving at different rates.
“The rapid mutation of the centromeric regions of the genome, along with their various mutation rates, has led to their diverse structure and organization,” Logsdon noted. It was surprising to learn, the scientists said, that such vital areas of the genome were subject to swift changes, because, in general, critical functions tend to be genetically conserved.
The scientists plan to expand these initial efforts by developing more genetic maps of centromeres in diverse human genomes and across various organs and tissues, and to perform multigeneration family studies of centromere sequences. The complete sequence of centromeres from other nonhuman primates will provide a better model of the evolutionary forces shaping these regions.
Peering into the future, Logsdon and her team hope someday to apply their findings on centromeres to the design and engineering of customized human artificial chromosomes to transform medical science. Several years ago, Logsdon and her mentors worked on efforts to develop human artificial chromosomes that bypass centromeric DNA, which had then posed a constraint to mammalian synthetic genome research. Logsdon and others recently published another study last week in Science, which showed that enlarging the artificial chromosome DNA vector allowed for efficient formation of human artificial chromosomes in cells.
The centromere studies reported today were supported by funding from the National Institutes of Health National Human Genome Research Institute (R01 HG010169); National Institute of General Medical Sciences (K99 GM147352 ); National Cancer Institute ( R01 CA266339); Intramural Research Program of the National Human Genome Research Institute; Shanghai Jiao Tong University 2030 Program (WH510363001-7); Center for Integration in Science of the Ministry of Aliyah, Israel; and the Howard Hughes Medical Institute. This work utilized the computational resources of the National Institutes of Health High Performance Computing Biowulf Linux cluster.
For details about UW Medicine, please see our About page.